Scientists found for the first time a natural protein that follows a mathematical pattern of self-similarity
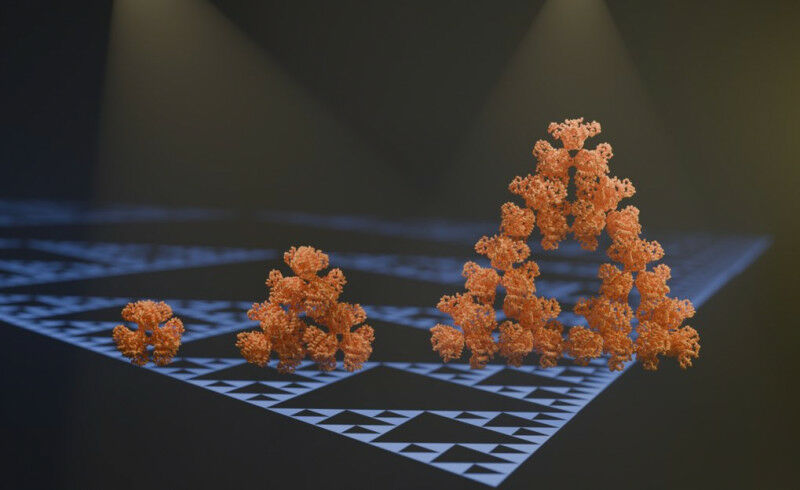

An international team of researchers led by groups from the Max Planck Institute in Marburg and the Philipps University in Marburg has stumbled upon the first regular molecular fractal in nature. They discovered a microbial enzyme - citrate synthase from a cyanobacterium - that spontaneously assembles into a pattern known as the Sierpinski triangle. Electron microscopy and evolutionary biochemistry studies indicate that this fractal may represent an evolutionary accident.
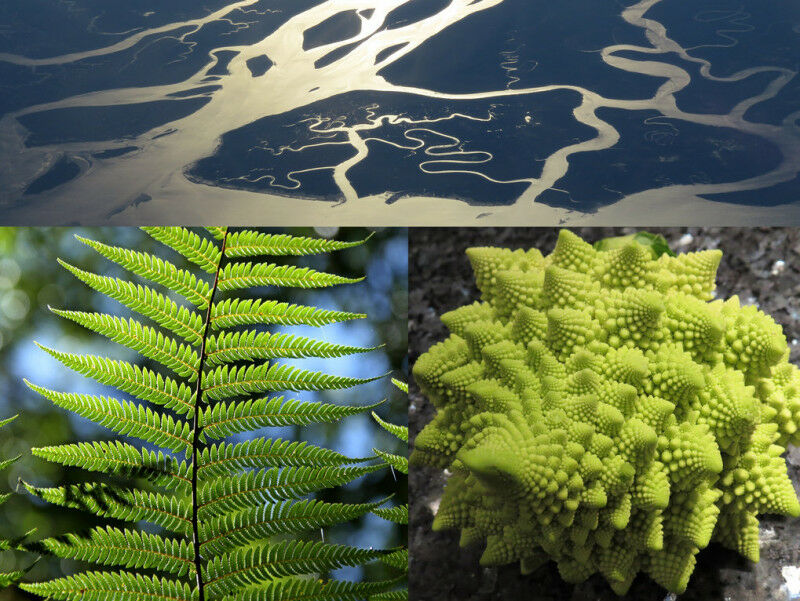
Snowflakes, fern leaves, romanesco cauliflower heads: many structures in nature have a certain regularity. Their individual parts resemble the shape of the whole structure. Such shapes, which repeat from the largest to the smallest, are called fractals. But regular fractals that match almost exactly across scales, as in the examples above, are very rare in nature.
Molecules also have a certain regularity. But if you look at them from a great distance, you can no longer see any signs of this. Then you see smooth matter whose features no longer match those of the individual molecules. The degree of fine structure we see depends on our magnification - in contrast to fractals, where self-similarity persists at all scales. In fact, regular fractals at the molecular level are completely unknown in nature.
This is somewhat surprising. After all, molecules can assemble themselves into all sorts of wonderful shapes. Scientists have extensive catalogues of self-assembled complex molecular structures. However, there has never been a regular fractal among them. It turns out that almost all regular-looking self-assemblies lead to the kind of regularity that becomes smooth on large scales.
An international team of researchers led by groups from the Max Planck Institute in Marburg and Phillips University in Marburg has now discovered the first regular molecular fractal in nature. They discovered a microbial enzyme - citrate synthase from a cyanobacterium - that spontaneously assembles into a regular fractal pattern known as the Sierpinski triangle. This is an infinitely repeating series of triangles made up of smaller triangles. ’We stumbled on this structure completely by accident and almost couldn’t believe what we saw when we first took images of it using an electron microscope’ says first author Franziska Sendker. ’The protein makes these beautiful triangles and as the fractal grows, we see these larger and larger triangular voids in the middle of them, which is totally unlike any protein assembly we’ve ever seen before’ she continues.
How did this unusual exception emerge? What distinguishes the enzyme from all’others, causing it to form a fractal shape? Teaming up with structural biologist at the university of Marburg, the team eventually managed to determine the molecular structure of this assembly using electron microscopy, which illuminated how it achieves its fractal geometry. ’This was one of the harder, but also more fascinating structures I have solved in my career’ says Jan Schuller, whose group helped determine the structure. ’The problem with determining the structure of a fractal is that our image averaging techniques kept getting confused by the fact that the smaller triangles can be substructures of larger triangles. The algorithm kept homing in on these smaller triangles instead of seeing the larger structures they were part of’ he explains.
Asymmetry leads to fractal formation

With the structure in hand, it became clear how exactly this protein manages to assemble into a fractal: Normally, when proteins self-assemble, the pattern is highly symmetrical: each individual protein chain adopts the same arrangement relative to its neighbours. Such symmetrical interactions always lead to patterns that become smooth on large scales. The key to the fractal protein was that its assembly violated this rule of symmetry. Different protein chains made slightly different interactions at different positions in the fractal. This was the basis for forming the Sierpinski triangle, with its large internal voids, rather than a regular lattice of molecules.
Does this bizarre assembly do anything useful? ’Self-assembly is often used by evolution to regulate enzymes, but in this case the cyanobacterium that this enzyme is found in does not seem to care much whether or not its citrate synthase can assemble into a fractal’, says evolutionary biologist Georg Hochberg, one of the senior authors of the study. When the team genetically manipulated the bacterium to prevent its citrate synthase from assembling into the fractal triangles, the cells grew just as well under a variety of conditions. ’This prompted us to wonder whether this might just be a harmless accident of evolution. Such accidents can happen when the structure in question isn’t too difficult to construct’.
Replaying evolution in the lab
To test their theory, the team recreated the evolutionary development of the fractal arrangement in the laboratory. To do this, they used a statistical method to back-calculate the protein sequence of the fractal protein as it was millions of years ago. By then producing these ancient proteins biochemically they were able to show that the arrangement arose quite suddenly through a very small number of mutations and was then immediately lost again in several cyanobacterial lineages, so that it only remained intact in this single bacterial species. ’Although we can never be totally sure of the reasons why things happened in the past, this particular case does have all the trappings of a seemingly complex biological structure that just popped into existence for no good reason at all because it was simply very easy to evolve’ says Hochberg.
The fact that something so complex-looking as a molecular fractal could emerge so easily in evolution suggests that more surprises and much beauty may still lie hidden in so far undiscovered molecular assemblies of many biomolecules.
Sendker, F.L.; Lo , J.K.; Heimerl, T.; Bohn, S.; Persson, L-J.; Mais, C.N.; Sadowska, W.; Paczia, N.; Nußbaum, E.; del Carmen Sánchez Olmos, M.; Forchhammer. K.; Schindler, D.; Erb, T.J.; Benesch, J.L.P.; Marklund, E.; Bange, G.; Schuller, J.M.; Hochberg, G.K.A.